
Made terminator features directional in the common features database Added clickable "contact support" links to messages that pop up in various places Enhanced enzyme, feature, and primer label layout in Map view for linear sequences

Added an indicator in the selection bar to show "DNA" or "RNA" as the sequence type
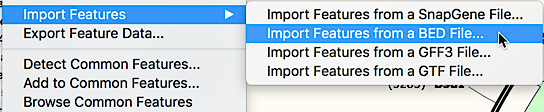
EXPORT GTF FROM SNAPGENE VIEWER PDF
Enabled printing to PDF on macOS when no printers are installed Added support for /protein_id qualifiers for mat_peptide features Enhanced the response to mousing over a primer so that the binding site is now highlighted in gray Enhanced the visualization of mutagenesis simulations and mutagenic primers Enabled export of alignments to rich text format. Extended the range for the minimum required 3' match for primers from 25 to 35 bases Added DNA Ladders from Ecogen, TIANGEN, and Vazyme geneious nucleotide and protein sequences and alignments Enabled viewing and printing chromatogram files in a wrapped multi-line format Added support for annotating primers that anneal at the 5’ end but not the 3’ end Added support for filling in DNA ends of annealed oligos to create a double-stranded DNA sequence Enabled gaps to be displayed every 10th base for protein sequences, and every 10th or 3rd base for nucleotide sequences Enabled viewing files of all types simultaneously in a Collection using a new "All Files" area Enhanced History view with a Text format tab, and with an option to show the entire history Added support for viewing features in multiple sequence alignments

